How to Investigate those Pesky Serologic Weak D Samples

When typing a patient for the RhD antigen, hospitals have been forced to convert a non-binary result into a positive or negative for purposes of entry into the electronic health record as well as for selection of blood products including Rh immune globulin (RhIG). RhD is not unique amongst blood group antigens in demonstrating a property by which, in some individuals, the antigen phenotype is weakly positive. It may also type positive with one reagent or methodology used for antigen typing while typing negative with another. Since alloimmunization to RhD is clinically significant, both in the context of pregnancy and transfusion, it is critical to manage patients with serologic weak D or RhD typing discrepancies appropriately.
It has been known for more than 30 years that the RHD gene is the source of much genetic variation that leads to variant antigens. The RhD antigen is complex and is comprised of many epitopes, the portions of the antigen that are bound by antibodies. Panels of monoclonal anti-D reagents can be used to show epitope loss. These antigens are typically categorized as weak, partial or DEL. The term “weak” when initially applied to RhD variants, was used to indicate that though the number of antigen copies present on the red blood cell surface was likely lower than normal; however, the epitopes that are detected with anti-D reagents were thought to be intact. The term ‘partial” when applied to RhD variants indicates that one or more of the epitopes has been destroyed or altered. The DEL phenotype describes red cells on which the RhD antigen (which may have been altered or lost epitopes), can only be detected by use of adsorption-elution techniques.
Since establishing this naming convention, some weak D types have been associated with allo-anti-D formation. Also, the RhD antigen in DEL phenotypes may or may not have all epitopes intact. Addtionally, some RHD alleles discovered by molecular approaches with no available information about alloimmunization were classified as weak or partial based on the location of the amino acid changes and the predicted impact on epitopes. The use of the term D variant has been suggested to describe any RhD antigen different from that encoded by the reference RHD allele, without requirement to have structural or clinical data for classification.
Prior to 2015, a phenotype was considered “weak D” if testing yielded a negative result at immediate spin but a positive result with the indirect antiglobulin test (IAT). This phenotype had been referred to as DU positive, but this term is no longer utilized. For the last 50 years or so, women who typed weakly positive with anti-D reagents were managed as RhD+. However, over the last 25 years, we have gained a better understanding of the genetic backgrounds of individuals in whom RhD type is not equivocal, as well as the clinical implications.
A survey of hospitals conducted in 2012 by the College of American Pathologists that generated responses from 3100 hospitals showed a lack of standardization for handling patients with serologic weak D.
In 2015, an interorganizational team representing AABB, CAP, ABC, ARC, ACOG published a commentary redefining a “serologic weak D” phenotype result with a strength of reactivity less than or equal to 2+, without specifying methodology or reagent type, or a subject with a discrepant type, which could involve current vs historic type.
This article summarized decades of research and laboratory findings and provided a workflow for when a hospital blood bank should consider using RHD genotyping to resolve a serologic weak D type. The approach is designed to precisely identify patients at risk of RhD alloimmunization and therefore candidates for Rh immunoprophylaxis or D negative red cell units for transfusion. A financial analysis indicated that this approach could be cost saving, though the greater benefit would arise from better portability of personal health information and/or a national electronic health record.
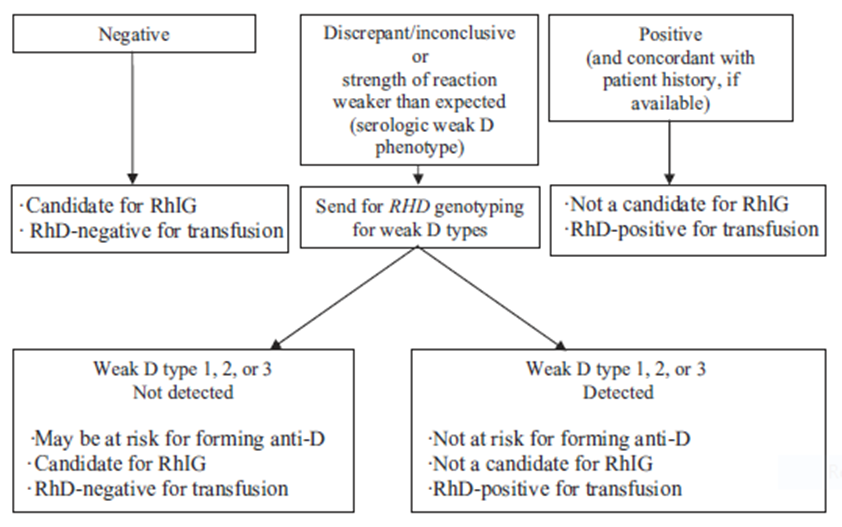
Figure 1. Algorithm for resolving serologic weak D phenotype test results by RHD genotyping to determine candidacy for RhIG and RhD type for transfusion
A joint statement by AABB, American Red Cross, ABC, CAP (https://www.cap.org/) later that year recommended implementation of the workflow in women of child-bearing age. Since then, many hospitals have been unsure of how to operationalize this recommendation. One large urban hospital evaluated three criteria and showed (in collaboration with the American Red Cross National Molecular Laboratory (NML)) that two provided excellent specificity for identifying samples that express RhD variants. Another hospital recently published their approach (also in collaboration with the American Red Cross NML) focused on obstetrics patients. Both studies demonstrate that discordant results with two test methodologies or anti-D reagents can effectively identify patients expressing D variants and that RHD genotyping can differentiate these patients into those at risk and those not at risk for allo-anti-D formation.
These studies highlight two important findings. First, subjects who express partial D can present with a serologic weak D phenotype. This finding should give those managing hospital blood banks pause if their policy would classify patients who type weakly with anti-D reagents as RhD+ since some of these patients may be at risk of forming allo-anti-D. Second, automated antigen typing instruments can result partial D subjects as D+ such that their risk of alloimmunization may be undetected. This also should concern blood bankers, since this finding would suggest that some patients who may be at risk of allo-anti-D formation are going undetected. This is especially concerning for women of child-bearing age and especially in women of African descent, in whom the partial D phenotype is most common.
In 2020, the interorganizational team reconvened to recommend that blood banks consider a lab value of “serologic weak D” an incomplete result. They recommend that the sample be tested, typically at a molecular reference laboratory, to determine alloimmunization risk. Specifically, RHD genotyping is used determine if the subject expresses a D variant, and if so, the identification of the variant is used to make an assessment of alloimmunization risk that can aid the clinical team regarding transfusion and immunoprophylaxis candidacy.
References
- Identifying obstetrics patients in whom RHD genotyping can be used to assess risk of D alloimmunization TN Horn et al Immunohematology 2020. 36 (4), 146-151
- It’s Time to Phase Out” Serologic Weak D Phenotype” and Resolve D Types With RHD Genotyping Including Weak D Type 4 WA Flegel et al Transfusion 2020. 60 (4), 855-859
- Experience with RHD* weak D type 4.0 in the USA CM Westhoff et al Blood Transfusion 2019. 17 (2), 91
- Strategies to identify candidates for D variant genotyping X Luo et al Blood Transfusion 2018. 16 (3), 293
- Financial implications of RHD genotyping of pregnant women with a serologic weak D phenotype S Kacker et al Transfusion 2015. 55 (9), 2095-2103
- It’s time to phase-in RHD genotyping for patients with a serological weak D phenotype
- SG Sandler et al Transfusion 2015. 55 (3), 680-689
- Serological weak D phenotypes: a review and guidance for interpreting the RhD blood type using the RHD genotype. SG Sandler et al Br J Haematol. 2017 Oct;179(1):10-19.
- Policies and procedures related to testing for weak D phenotypes and administration of Rh immune globulin: results and recommendations related to supplemental questions in the Comprehensive Transfusion Medicine survey of the College of American Pathologists S Gerald Sandler et al Arch Pathol Lab Med. 2014 May;138(5):620-5.
- Variants of RhD – current testing and clinical consequences G Daniels Br J Haematol 2013 161:461-470.
- How do I manage Rh typing in obstetric patients? Haspel RL & Westhoff CM. Transfusion 2015,55:470-4.
- How I manage donors and patients with a weak D phenotype. Flegel, WA, Curr Opin Hematol 2006,13:476–483.
- Tentative model for the mapping of D epitopes on the RhD polypeptide J P Cartron et al. Transfus Clin Biol. 1996;3(6):497-503.
Additional Resources
- AABB Joint Statement on Phasing-In RHD Genotyping for Pregnant Women and Other Females of Childbearing Potential with a Serologic Weak D Phenotype: https://www.aabb.org/docs/default-source/default-document-library/positions/statement150722.pdf?sfvrsn=f4f44c30_6
- American Red Cross Immunohematology Reference Lab Testing (IRL) Website: https://www.redcrossblood.org/biomedical-services/blood-diagnostic-testing/immunohematology-reference-lab-testing.html
- American Red Cross Molecular Testing Website: https://www.redcrossblood.org/biomedical-services/blood-diagnostic-testing/molecular-testing.html
- American Red Cross SUCCESS Webinar: All Things Rh and The Rh Blood Group System, accessible at SuccessEducation.Redcross.org.
Written By
Margaret A Keller, PhD
Executive of the Red Cross National Laboratories and the American Rare Donor Program. She is also the Editor-in-chief of Immunohematology, Journal of Blood Group Serology and Molecular Genetics. Dr. Keller has over 25 years of experience in molecular genetics and for the last decade has been involved in using molecular methods to identify rare and antigen negative red cell and platelet donors as well as to assess patient risk of alloimmunization associated with transfusion or pregnancy. Dr. Keller is interested in genetic variation in the RH locus and using RH allele information for selection of blood products for Rh alloimmunized patients. Dr. Keller is adjunct associate professor at Thomas Jefferson University where she is involved in training of Transfusion Medicine fellows as well as students in Jefferson genetic counselor program.